Scientific Reports published results of a new Mycobacterium tuberculosis study led by Igor Mokrousov (St. Petersburg Pasteur Institute) that joined efforts of scientists from Russian regions, Europe, Asia and South America. The authors have developed a molecular method for rapid detection of strains of the emerging genotype Mycobacterium tuberculosis Beijing 1071-32, which is characterized by multidrug resistance. The study was supported by the Russian Science Foundation, grant 19-14-00013.
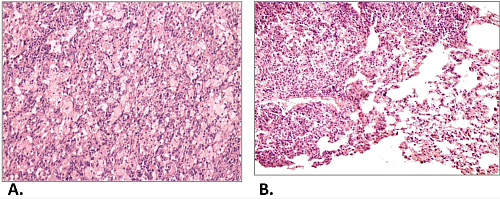
M. tuberculosis is a sufficiently diverse species and clinical significance was associated with some of its genetic families or clonal clusters. In-depth analysis of medically important genotypes allows for detection of their specific markers associated with pathobiologically significant features such as drug resistance, hypervirulence, or transmissibility. Whole-genome and next-generation sequencing (WGS/NGS) is a comprehensive method of pathogen analysis. This key to open all (or many) doors is becoming less expensive and more available. Nonetheless it is unrealistic to implement NGS for routine TB diagnostics and surveillance in the high burden countries such as Russia where 72000 new cases were reported in 2019. This highlights the vital interest in the development of rapid, low-cost, and high-performance methods for detection of the particular epidemic strains.
Russian members of the research team have previously discovered emerging and highly resistant genetic clusters of M. tuberculosis in the Omsk region of Western Siberia. These strains belonged to the rarely seen in Russia ancient sublineage of the Beijing genotype. Some strains were shown to be highly lethal and hypervirulent in the mouse model experiment. Most of the isolates belonged to the Beijing 1071-32 cluster and all were multidrug resistant (MDR).
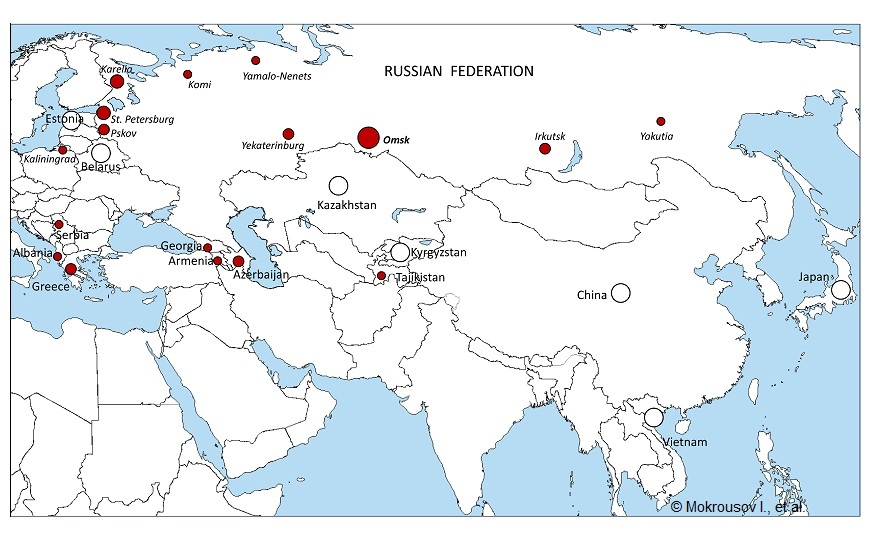
In the new study, based on the phylogenetic analysis of the genomic data, authors identified three cluster-specific synonymous SNPs in the genes Rv0144, Rv0373c, and Rv0334 and developed and validated the real-time PCR assay for their detection. Analysis of the genetically and geographically diverse collection of ~2400 M. tuberculosis isolates sampled in 1996 to 2020 (European and Asian parts of Russia, former Soviet Union countries, Albania, Greece, China, Vietnam, Japan and Brazil), confirmed 100% specificity and sensitivity of this assay.
All Beijing 1071-32 isolates carried a characteristic signature of six mutations that confer resistance to the four first-line antibiotics. Intriguingly, this combination includes the most frequent and efficient mutations (rpoB450, katG315, rpsL43), rare mutation (embB497), and compensatory mutations (rpoC485, katG335). The latter are supposed to restore the reduced strain fitness caused by previously acquired resistance mutations. The epistatic interaction of all these mutations could have influenced the spread of this genetic cluster.
In addition to the expected presence of the Beijing 1071-32 in Siberia and the European part of Russia, these strains were found in Central Asia, Transcaucasia, and also, quite unexpectedly, in the Balkan countries - Albania, Greece, Serbia. It is unknown whether the latter reflects already the local Balkan circulation of this strain or independent events of its introduction from the countries of the former USSR.
DNA analysis of samples from Albania and Greece confirmed that they harbored all 9 targeted mutations, i.e. 3 neutral SNPs and 6 resistance mutations. Greek isolates were recovered one from an immigrant from Georgia, the other two from local patients. Since the Beijing genotype is extremely rare in Albania, its isolates probably came to this country from the neighboring Greece, which is characterized by a noticeable presence of immigrants (and MDR strains) from the former USSR. A large European multicenter study of drug resistant TB in the EU ten years ago identified the Beijing 1071-32 isolates and designated them as European resistant cluster ECDC_10. This highlights the presence of this genetic cluster in the EU and the relevance of its tracing on a more global scale not limited to Russia and other Post-Soviet states.
According to Dr Igor Mokrousov, the Russian (presumably Siberian) origin of the Beijing 1071-32 cluster is the most likely scenario, since this cluster includes isolates from remote parts of Russia, with the highest prevalence in Omsk, Siberia.
“The circumstances of its origin remain unknown,” says Mokrousov. “The notorious Russian epidemic strain Beijing B0/W148 probably originated in Siberia in the 1950s and spread in the prisons later on. It is possible that the Beijing 1071-32 also initially originated and disseminated within the Soviet penitentiary system in the 1970s”.
rpoB and rpoC mutations found in the Beijing 1071-32 strain are known to cause resistance to rifampicine. Interestingly, phylogenomic analysis suggested the time of origin of this strain in the 1970s – and this was time when rifampicine was first included into anti-TB chemotherapy. This scenario resembles the situation in Portugal and South Africa, where endemic strains of the Latin American Mediterranean genetic family evolved towards multidrug-resistance in the 1970s.
Enigmatically, all geographically distant isolates of this Beijing 1071-32 cluster have the same set of six resistance mutations. No intermediate strains with some, but not all these mutations were found to date.
“Speculatively, the acquisition of new mutations made the strain more adaptable compared to intermediate variants that subsequently disappeared from the population,” says Igor Mokrousov.
Citation: Mokrousov I. et al. Practical approach to detection and surveillance of emerging highly resistant Mycobacterium tuberculosis Beijing 1071-32-cluster. Scientific Reports 11, 21392 (2021). DOI 10.1038/s41598-021-00890-7. https://www.nature.com/articles/s41598-021-00890-7